What is Bayesian Optimization?
Bayesian optimization is a method for global optimization of expensive-to-evaluate black-box functions. It targets problems of the form:
\[ \mathbf{x}^* = \arg\min_{\mathbf{x} \in \mathcal{X}} f(\mathbf{x}) \tag{1} \]where \(f\) has no analytical gradient and each evaluation incurs a significant cost in time or resources. Common applications include hyperparameter tuning for machine learning models, experimental design, and materials discovery.
Bayesian optimization consists of two components:
- Surrogate model: A probabilistic model that approximates the objective function (typically a Gaussian process)
- Acquisition function: A criterion that determines which point to evaluate next
By building a surrogate from a small set of observations and sequentially selecting points that maximize the acquisition function, Bayesian optimization explores the search space efficiently.
Gaussian Process Regression
Kernel Functions
A Gaussian process (GP) defines a prior distribution over functions. A GP is fully specified by a mean function \(m(\mathbf{x})\) and a kernel function \(k(\mathbf{x}, \mathbf{x}')\):
\[ f(\mathbf{x}) \sim \mathcal{GP}(m(\mathbf{x}), k(\mathbf{x}, \mathbf{x}')) \tag{2} \]The most widely used kernel is the RBF (Radial Basis Function) kernel, also known as the squared exponential kernel:
\[ k(\mathbf{x}, \mathbf{x}') = \sigma_f^2 \exp\left(-\frac{\|\mathbf{x} - \mathbf{x}'\|^2}{2l^2}\right) \tag{3} \]where \(\sigma_f^2\) is the output variance and \(l\) is the length-scale parameter. Larger values of \(l\) produce smoother functions.
Posterior Distribution
Given \(n\) observations \(\mathcal{D} = \{(\mathbf{x}_i, y_i)\}_{i=1}^n\) with observation noise \(\sigma_n^2\), the predictive distribution at a new input \(\mathbf{x}_*\) is:
\[ \mu(\mathbf{x}_*) = \mathbf{k}_*^T (\mathbf{K} + \sigma_n^2 \mathbf{I})^{-1} \mathbf{y} \tag{4} \]\[ \sigma^2(\mathbf{x}_*) = k(\mathbf{x}_*, \mathbf{x}_*) - \mathbf{k}_*^T (\mathbf{K} + \sigma_n^2 \mathbf{I})^{-1} \mathbf{k}_* \tag{5} \]where:
- \(\mathbf{K}\) is the \(n \times n\) kernel matrix with \(K_{ij} = k(\mathbf{x}_i, \mathbf{x}_j)\)
- \(\mathbf{k}_* = [k(\mathbf{x}_1, \mathbf{x}_*), \ldots, k(\mathbf{x}_n, \mathbf{x}_*)]^T\)
- \(\mathbf{y} = [y_1, \ldots, y_n]^T\)
Equation (4) gives the predictive mean, and Equation (5) gives the predictive uncertainty. The variance \(\sigma^2\) shrinks in regions dense with data and grows in regions with sparse observations.
Acquisition Functions
Acquisition functions decide where to sample next. They balance exploitation (regions with low predicted mean) and exploration (regions with high uncertainty).
Expected Improvement (EI)
EI maximizes the expected improvement over the current best observed value \(f(\mathbf{x}^+)\):
\[ \text{EI}(\mathbf{x}) = \mathbb{E}[\max(f(\mathbf{x}^+) - f(\mathbf{x}), 0)] \tag{6} \]Under the GP predictive distribution, EI admits a closed-form expression:
\[ \text{EI}(\mathbf{x}) = (\mu(\mathbf{x}^+) - \mu(\mathbf{x}) - \xi) \Phi(Z) + \sigma(\mathbf{x}) \phi(Z) \tag{7} \]\[ Z = \frac{\mu(\mathbf{x}^+) - \mu(\mathbf{x}) - \xi}{\sigma(\mathbf{x})} \tag{8} \]where \(\Phi\) and \(\phi\) are the CDF and PDF of the standard normal distribution, and \(\xi \geq 0\) is a parameter that encourages exploration. For minimization, \(f(\mathbf{x}^+)\) is the current minimum observation.
Upper Confidence Bound (UCB)
For minimization, we use the Lower Confidence Bound variant:
\[ \text{UCB}(\mathbf{x}) = \mu(\mathbf{x}) - \kappa \sigma(\mathbf{x}) \tag{9} \]The parameter \(\kappa > 0\) controls the exploration–exploitation trade-off. Larger \(\kappa\) favors exploration.
Probability of Improvement (PI)
PI maximizes the probability of improving upon the current best value:
\[ \text{PI}(\mathbf{x}) = \Phi\left(\frac{\mu(\mathbf{x}^+) - \mu(\mathbf{x}) - \xi}{\sigma(\mathbf{x})}\right) \tag{10} \]PI is simple to compute but ignores the magnitude of improvement, which can lead to premature convergence to local optima.
The figure below compares how the three acquisition functions suggest different exploration strategies for the same GP surrogate.
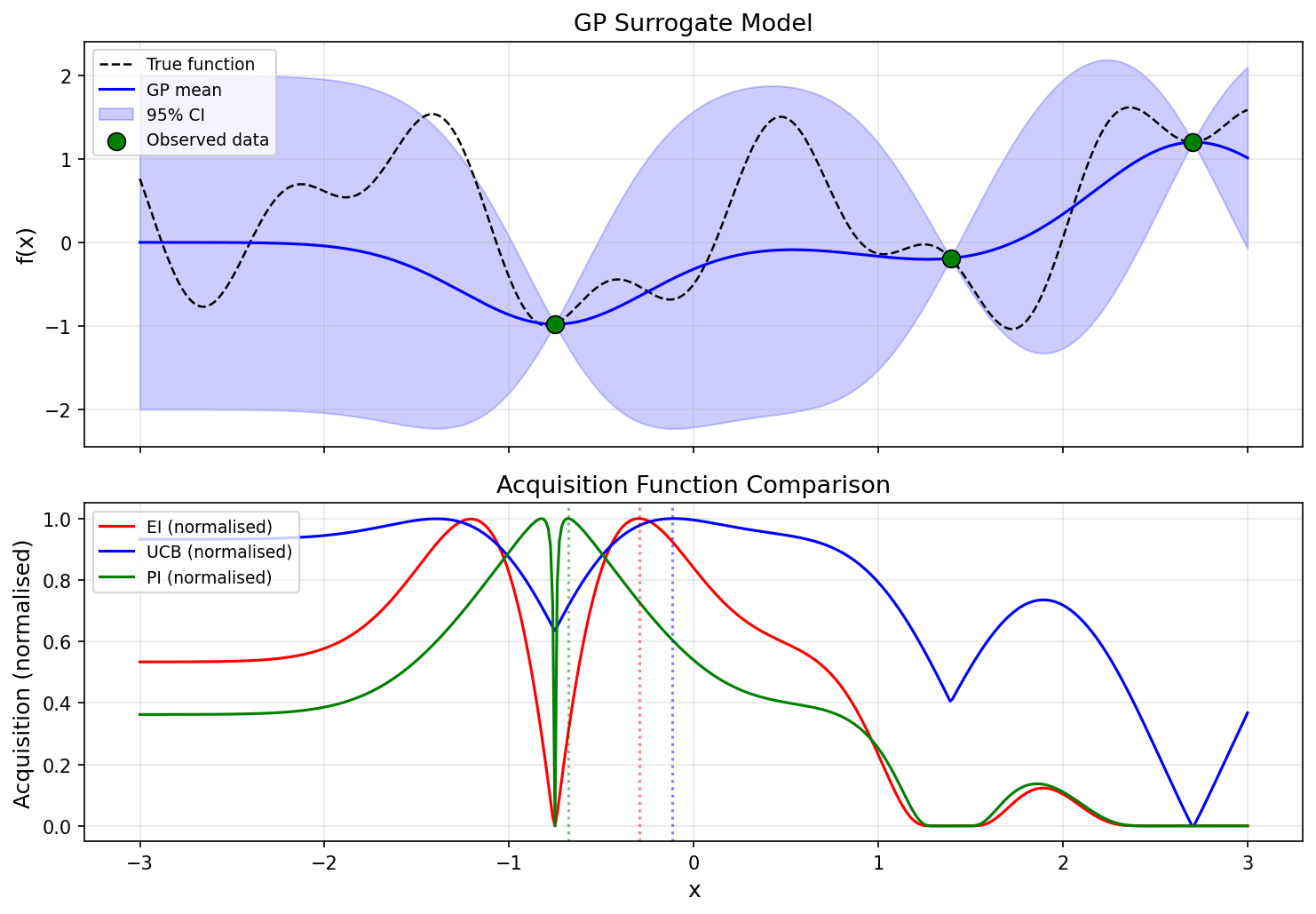
Python Implementation
Gaussian Process Class
We implement GP regression from scratch using NumPy and SciPy.
import numpy as np
from scipy.stats import norm
class GaussianProcess:
def __init__(self, length_scale=1.0, signal_var=1.0, noise_var=1e-6):
self.l = length_scale
self.sf2 = signal_var
self.sn2 = noise_var
self.X_train = None
self.y_train = None
self.K_inv = None
def kernel(self, X1, X2):
"""RBF kernel (Eq. 3)"""
dist_sq = np.sum(X1**2, axis=1, keepdims=True) \
- 2.0 * X1 @ X2.T \
+ np.sum(X2**2, axis=1)
return self.sf2 * np.exp(-0.5 * dist_sq / self.l**2)
def fit(self, X, y):
"""Fit the GP to observed data"""
self.X_train = X.copy()
self.y_train = y.copy()
K = self.kernel(X, X) + self.sn2 * np.eye(len(X))
self.K_inv = np.linalg.inv(K)
def predict(self, X_new):
"""Predictive mean and variance at new points (Eqs. 4, 5)"""
k_star = self.kernel(self.X_train, X_new)
mu = k_star.T @ self.K_inv @ self.y_train
var = self.sf2 - np.diag(k_star.T @ self.K_inv @ k_star)
var = np.maximum(var, 1e-10)
return mu, var
Acquisition Functions
def expected_improvement(mu, var, y_best, xi=0.01):
"""Expected Improvement (Eqs. 7, 8)"""
sigma = np.sqrt(var)
with np.errstate(divide="ignore", invalid="ignore"):
Z = (y_best - mu - xi) / sigma
ei = (y_best - mu - xi) * norm.cdf(Z) + sigma * norm.pdf(Z)
ei[sigma < 1e-10] = 0.0
return ei
def upper_confidence_bound(mu, var, kappa=2.0):
"""UCB for minimization (LCB) (Eq. 9)"""
sigma = np.sqrt(var)
return mu - kappa * sigma
def probability_of_improvement(mu, var, y_best, xi=0.01):
"""Probability of Improvement (Eq. 10)"""
sigma = np.sqrt(var)
with np.errstate(divide="ignore", invalid="ignore"):
Z = (y_best - mu - xi) / sigma
pi = norm.cdf(Z)
pi[sigma < 1e-10] = 0.0
return pi
Bayesian Optimization Loop
def bayesian_optimization(f, bounds, n_init=3, n_iter=15,
acq_func="ei"):
"""Main Bayesian optimization loop"""
# Random initial samples
X_init = np.random.uniform(bounds[0], bounds[1],
size=(n_init, 1))
y_init = np.array([f(x) for x in X_init]).reshape(-1)
gp = GaussianProcess(length_scale=0.5, signal_var=1.0,
noise_var=1e-6)
X_sample = X_init.copy()
y_sample = y_init.copy()
# Optimization loop
for i in range(n_iter):
gp.fit(X_sample, y_sample)
# Evaluate acquisition function over candidate points
X_cand = np.linspace(bounds[0], bounds[1], 500).reshape(-1, 1)
mu, var = gp.predict(X_cand)
y_best = np.min(y_sample)
if acq_func == "ei":
acq = expected_improvement(mu, var, y_best)
x_next = X_cand[np.argmax(acq)]
elif acq_func == "ucb":
acq = upper_confidence_bound(mu, var)
x_next = X_cand[np.argmin(acq)]
elif acq_func == "pi":
acq = probability_of_improvement(mu, var, y_best)
x_next = X_cand[np.argmax(acq)]
# Evaluate and add the new point
y_next = f(x_next.reshape(1, -1))
X_sample = np.vstack([X_sample, x_next.reshape(1, -1)])
y_sample = np.append(y_sample, y_next)
return X_sample, y_sample, gp
1D Optimization Demo
We use a non-convex test function with multiple local minima to demonstrate Bayesian optimization.
import matplotlib.pyplot as plt
def test_function(x):
"""Test function with multiple local minima"""
return np.sin(3 * x) + x**2 * 0.1 - 0.5 * np.cos(7 * x)
np.random.seed(42)
bounds = (-3.0, 3.0)
X_sample, y_sample, gp = bayesian_optimization(
test_function, bounds, n_init=3, n_iter=12, acq_func="ei"
)
# Plot the results
X_plot = np.linspace(bounds[0], bounds[1], 300).reshape(-1, 1)
y_true = np.array([test_function(x) for x in X_plot])
mu, var = gp.predict(X_plot)
sigma = np.sqrt(var)
fig, axes = plt.subplots(2, 1, figsize=(10, 8))
# GP surrogate
axes[0].plot(X_plot, y_true, "k--", label="True function")
axes[0].plot(X_plot, mu, "b-", label="GP mean")
axes[0].fill_between(X_plot.ravel(), mu - 2*sigma, mu + 2*sigma,
alpha=0.2, color="blue", label="95% CI")
axes[0].scatter(X_sample[:3], y_sample[:3],
c="green", s=80, zorder=5, label="Initial points")
axes[0].scatter(X_sample[3:], y_sample[3:],
c="red", s=80, zorder=5, label="BO samples")
axes[0].set_xlabel("x")
axes[0].set_ylabel("f(x)")
axes[0].set_title("Bayesian Optimization with GP Surrogate")
axes[0].legend()
axes[0].grid(True)
# EI acquisition function
acq_ei = expected_improvement(mu, var, np.min(y_sample))
axes[1].plot(X_plot, acq_ei, "r-", label="EI")
axes[1].set_xlabel("x")
axes[1].set_ylabel("EI(x)")
axes[1].set_title("Expected Improvement")
axes[1].legend()
axes[1].grid(True)
plt.tight_layout()
plt.savefig("bayesian_optimization_result.png", dpi=150)
plt.show()
best_idx = np.argmin(y_sample)
print(f"Best x: {X_sample[best_idx].item():.4f}")
print(f"Best f(x): {y_sample[best_idx]:.4f}")
print(f"Total evaluations: {len(y_sample)}")
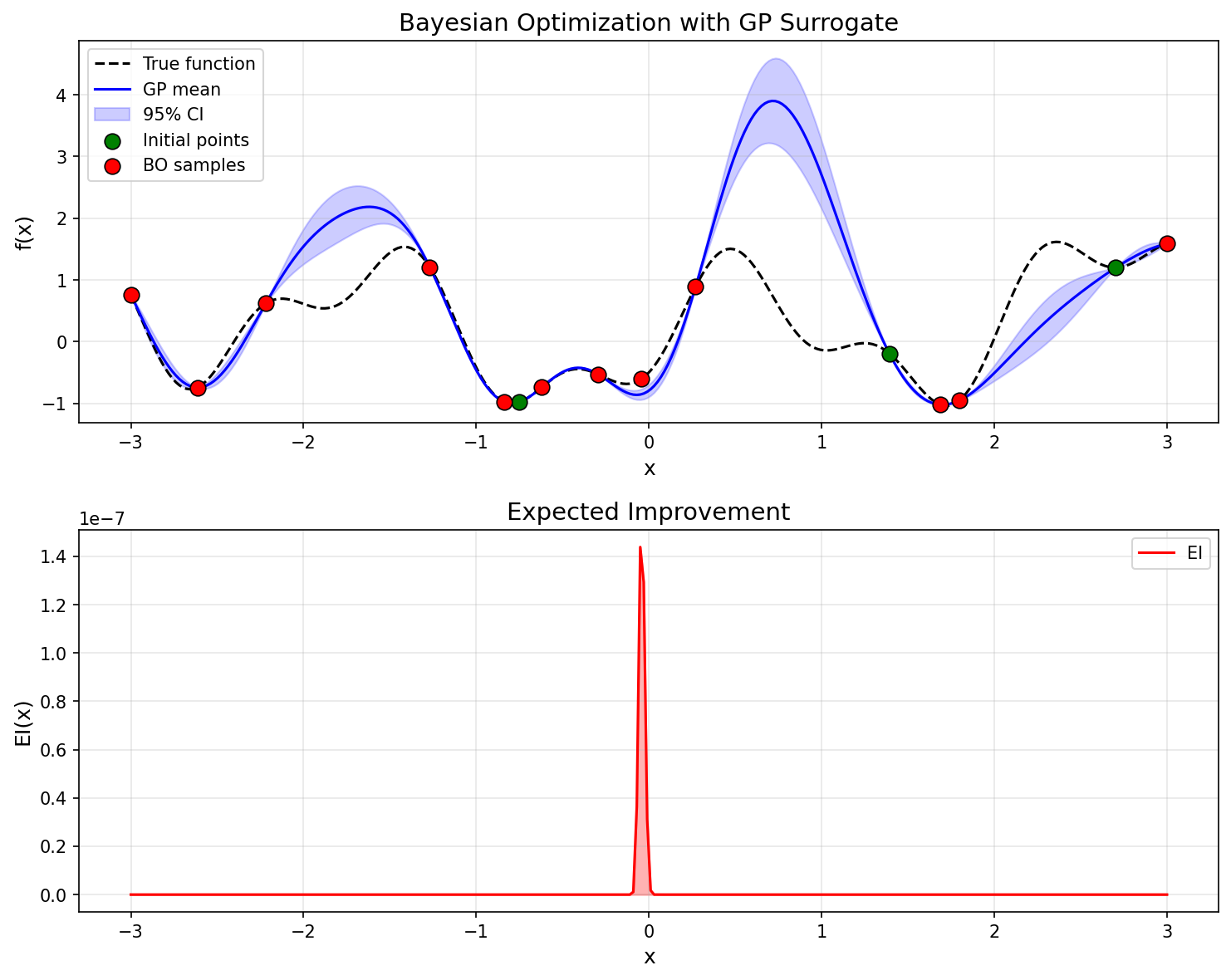
The GP predictive mean (blue line) converges toward the true function (dashed black line), while the confidence interval (blue band) narrows near observed data. The EI acquisition function takes large values in regions where uncertainty is high and improvement is expected, automatically balancing exploration and exploitation.
Comparison with CEM and MPPI
Bayesian optimization takes a fundamentally different approach from Cross-Entropy Method (CEM) and MPPI. The following table highlights the key differences.
| Bayesian Optimization | CEM | MPPI | |
|---|---|---|---|
| Evaluation budget | Very small (tens) | Moderate (hundreds–thousands) | Moderate (hundreds–thousands) |
| Parallelism | Sequential (batch extensions exist) | High | High |
| Gradient-free | Yes | Yes | Yes |
| Surrogate model | Yes (GP) | No | No |
| Exploration strategy | Acquisition function | Elite sample selection | Exponential weighting |
| Primary use case | Hyperparameter tuning | Combinatorial optimization, planning | Real-time control |
| Scalability | Low-dimensional (~20D) | Scales to high dimensions | Scales to high dimensions |
- Bayesian optimization excels when each evaluation is expensive. The surrogate model enables efficient search with few evaluations, though GP computation costs limit applicability to high-dimensional problems.
- CEM is a sample-based method that works well when many evaluations are feasible. It updates the sampling distribution through hard selection of elite samples.
- MPPI is also sample-based but uses soft exponential weighting to leverage information from all samples. It is well-suited for real-time control problems.
Related Articles
- Simulated Annealing: Theory and Python Implementation - Temperature-based stochastic search metaheuristic.
- Genetic Algorithms: Fundamentals and Python Implementation - Evolutionary computation-based black-box optimization.
- From SGD to Adam: Evolution of Gradient-Based Optimization - Gradient-based optimization; Bayesian optimization is used for hyperparameter tuning.
- Markov Chain Monte Carlo (MCMC): Metropolis-Hastings and Gibbs Sampling - Computational methods for Bayesian inference underlying Bayesian optimization.
References
- Rasmussen, C. E., & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press.
- Shahriari, B., et al. (2016). “Taking the Human Out of the Loop: A Review of Bayesian Optimization.” Proceedings of the IEEE, 104(1), 148-175.
- Snoek, J., Larochelle, H., & Adams, R. P. (2012). “Practical Bayesian Optimization of Machine Learning Algorithms.” NeurIPS 2012.
- Brochu, E., Cora, V. M., & de Freitas, N. (2010). “A Tutorial on Bayesian Optimization of Expensive Cost Functions.” arXiv:1012.2599.