What is a Particle Filter?
The particle filter (also known as the Sequential Monte Carlo method, SMC) is a nonlinear filtering technique that represents the posterior distribution of the state using a set of weighted samples called particles. Unlike Gaussian-approximation filters, it requires no distributional assumptions and can handle arbitrary nonlinear and non-Gaussian systems.
Gaussian-approximation filters such as the Cubature Kalman Filter (CKF) and the Unscented Transform-based Unscented Kalman Filter (UKF) degrade in accuracy when the posterior distribution is multimodal or the system exhibits strong nonlinearities. The particle filter remains effective in these situations.
The particle filter shares its Monte Carlo sampling foundation with the Cross-Entropy Method (CEM), but whereas CEM targets optimization problems, the particle filter targets sequential state estimation in time series.
Algorithm Overview
State-Space Model
Consider the following nonlinear state-space model:
\[ x_k = f(x_{k-1}) + w_{k-1}, \quad w_{k-1} \sim p_w \]\[ y_k = h(x_k) + v_k, \quad v_k \sim p_v \]where \(f\) is the state transition function, \(h\) is the observation function, and \(w_{k-1}\), \(v_k\) are the process and observation noise respectively. The noise distributions \(p_w, p_v\) need not be Gaussian.
Sequential Monte Carlo Framework
The particle filter approximates the posterior distribution \(p(x_k | y_{1:k})\) using \(N\) weighted particles \(\{x_k^{(i)}, w_k^{(i)}\}_{i=1}^{N}\). Each step consists of three phases.
1. Prediction: Propagate each particle through the state transition model:
\[ x_k^{(i)} \sim p(x_k | x_{k-1}^{(i)}) \]In practice, this is implemented as \(x_k^{(i)} = f(x_{k-1}^{(i)}) + w_{k-1}^{(i)}\) by adding process noise.
2. Weight Update: Compute the likelihood of the new observation \(y_k\) for each particle and update the weights:
\[ w_k^{(i)} \propto p(y_k | x_k^{(i)}) \]3. Normalization: Normalize the weights so they sum to one:
\[ \tilde{w}_k^{(i)} = \frac{w_k^{(i)}}{\sum_{j=1}^{N} w_k^{(j)}} \]The state estimate is then computed as a weighted average:
\[ \hat{x}_k = \sum_{i=1}^{N} \tilde{w}_k^{(i)} x_k^{(i)} \]Effective Sample Size and Degeneracy
A major challenge in particle filtering is weight degeneracy: over time, most particles acquire negligibly small weights while a few particles dominate. This degeneracy is quantified by the Effective Sample Size (ESS):
\[ N_{\text{eff}} = \frac{1}{\sum_{i=1}^{N} (\tilde{w}_k^{(i)})^2} \]\(N_{\text{eff}}\) ranges from 1 to \(N\). When all weights are equal (no degeneracy), \(N_{\text{eff}} = N\). Resampling is triggered when \(N_{\text{eff}}\) drops below a threshold, typically \(N/2\).
def effective_sample_size(weights):
"""Compute the effective sample size."""
return 1.0 / np.sum(weights ** 2)
Resampling Methods
Resampling reconstructs the particle set by duplicating high-weight particles and discarding low-weight ones. Below are three widely used methods.
Systematic Resampling
Systematic resampling generates a single random number \(u_0\) and places equally spaced points along the cumulative distribution function (CDF). It has \(O(N)\) complexity and low variance, making it the most commonly used method in practice.
def systematic_resampling(weights):
"""Systematic resampling."""
N = len(weights)
positions = (np.random.random() + np.arange(N)) / N
cumsum = np.cumsum(weights)
indices = np.zeros(N, dtype=int)
i, j = 0, 0
while i < N:
if positions[i] < cumsum[j]:
indices[i] = j
i += 1
else:
j += 1
return indices
Residual Resampling
Residual resampling first deterministically copies each particle \(\lfloor N \tilde{w}^{(i)} \rfloor\) times, then samples the remaining particles probabilistically from the residual weights. The deterministic component reduces variance compared to purely random methods.
def residual_resampling(weights):
"""Residual resampling."""
N = len(weights)
indices = []
# Deterministic copies
num_copies = (N * weights).astype(int)
for i in range(N):
indices.extend([i] * num_copies[i])
# Handle residuals
residual_weights = N * weights - num_copies
residual_weights /= residual_weights.sum()
num_remaining = N - len(indices)
if num_remaining > 0:
cumsum = np.cumsum(residual_weights)
# Apply systematic resampling to the residual part
positions = (np.random.random() + np.arange(num_remaining)) / num_remaining
i, j = 0, 0
while i < num_remaining:
if positions[i] < cumsum[j]:
indices.append(j)
i += 1
else:
j += 1
return np.array(indices)
Stratified Resampling
Stratified resampling divides the \([0, 1)\) interval into \(N\) equal strata and draws an independent uniform random number within each stratum to sample from the CDF. It is similar to systematic resampling but uses independent random numbers per stratum.
def stratified_resampling(weights):
"""Stratified resampling."""
N = len(weights)
positions = (np.random.random(N) + np.arange(N)) / N
cumsum = np.cumsum(weights)
indices = np.zeros(N, dtype=int)
i, j = 0, 0
while i < N:
if positions[i] < cumsum[j]:
indices[i] = j
i += 1
else:
j += 1
return indices
| Property | Systematic | Residual | Stratified |
|---|---|---|---|
| Complexity | O(N) | O(N) | O(N) |
| Variance | Low | Lowest | Low |
| Random draws | 1 | Remaining only | N |
| Deterministic component | No | Yes (floor copies) | No |
Nonlinear System Demo
Benchmark Model
To evaluate particle filter performance, we use the widely adopted Univariate Nonstationary Growth Model. This model exhibits strong nonlinearity that challenges Gaussian-approximation filters.
State transition model:
\[ x_k = \frac{x_{k-1}}{2} + \frac{25 x_{k-1}}{1 + x_{k-1}^2} + 8\cos(1.2k) + w_k, \quad w_k \sim \mathcal{N}(0, \sigma_w^2) \]Observation model:
\[ y_k = \frac{x_k^2}{20} + v_k, \quad v_k \sim \mathcal{N}(0, \sigma_v^2) \]A key feature of this model is that the observation function \(h(x) = x^2/20\) is an even function, so positive and negative state values are indistinguishable from observations alone. This can produce a bimodal posterior distribution.
Particle Filter Implementation
import numpy as np
import matplotlib.pyplot as plt
# ---- Model definition ----
sigma_w = np.sqrt(10.0) # process noise std
sigma_v = np.sqrt(1.0) # observation noise std
def state_transition(x, k):
"""State transition function."""
return x / 2.0 + 25.0 * x / (1.0 + x ** 2) + 8.0 * np.cos(1.2 * k)
def observation(x):
"""Observation function."""
return x ** 2 / 20.0
def log_likelihood(y, x):
"""Log-likelihood log p(y|x)."""
diff = y - observation(x)
return -0.5 * (diff ** 2) / (sigma_v ** 2)
# ---- Particle filter ----
def particle_filter(y_obs, N_particles, resample_fn, resample_threshold=0.5):
"""
Run the particle filter.
Parameters
----------
y_obs : array, observation sequence
N_particles : int, number of particles
resample_fn : callable, resampling function
resample_threshold : float, ESS threshold (ratio of N_particles)
Returns
-------
x_est : array, state estimate sequence
ess_history : array, ESS history
"""
T = len(y_obs)
threshold = resample_threshold * N_particles
# Initialize: sample from prior
particles = np.random.normal(0, np.sqrt(5.0), N_particles)
weights = np.ones(N_particles) / N_particles
x_est = np.zeros(T)
ess_history = np.zeros(T)
for k in range(T):
# Prediction: state transition + process noise
particles = state_transition(particles, k + 1) \
+ sigma_w * np.random.randn(N_particles)
# Weight update: compute in log domain for numerical stability
log_w = log_likelihood(y_obs[k], particles)
log_w -= np.max(log_w) # numerical stabilization
weights = np.exp(log_w)
weights /= np.sum(weights)
# State estimate
x_est[k] = np.sum(weights * particles)
# ESS computation
ess = effective_sample_size(weights)
ess_history[k] = ess
# Resampling (ESS threshold-based)
if ess < threshold:
indices = resample_fn(weights)
particles = particles[indices]
weights = np.ones(N_particles) / N_particles
return x_est, ess_history
Running the Simulation
np.random.seed(42)
T = 100 # number of time steps
N_particles = 500 # number of particles
# Generate true state and observations
x_true = np.zeros(T)
y_obs = np.zeros(T)
x_true[0] = 0.1 # initial state
for k in range(1, T):
x_true[k] = state_transition(x_true[k - 1], k) \
+ sigma_w * np.random.randn()
for k in range(T):
y_obs[k] = observation(x_true[k]) + sigma_v * np.random.randn()
# Run particle filter with three resampling methods
resamplers = {
"Systematic": systematic_resampling,
"Residual": residual_resampling,
"Stratified": stratified_resampling,
}
results = {}
for name, fn in resamplers.items():
np.random.seed(42) # same seed for fair comparison
x_est, ess_hist = particle_filter(y_obs, N_particles, fn)
rmse = np.sqrt(np.mean((x_true - x_est) ** 2))
results[name] = {"estimate": x_est, "ess": ess_hist, "rmse": rmse}
print(f"{name}: RMSE = {rmse:.4f}")
Visualization
fig, axes = plt.subplots(2, 1, figsize=(12, 8), sharex=True)
# State estimation comparison
axes[0].plot(x_true, "k-", linewidth=1.5, label="True state")
colors = {"Systematic": "tab:blue", "Residual": "tab:orange",
"Stratified": "tab:green"}
for name, res in results.items():
axes[0].plot(res["estimate"], "--", color=colors[name],
linewidth=1, label=f"{name} (RMSE={res['rmse']:.2f})")
axes[0].set_ylabel("State $x_k$")
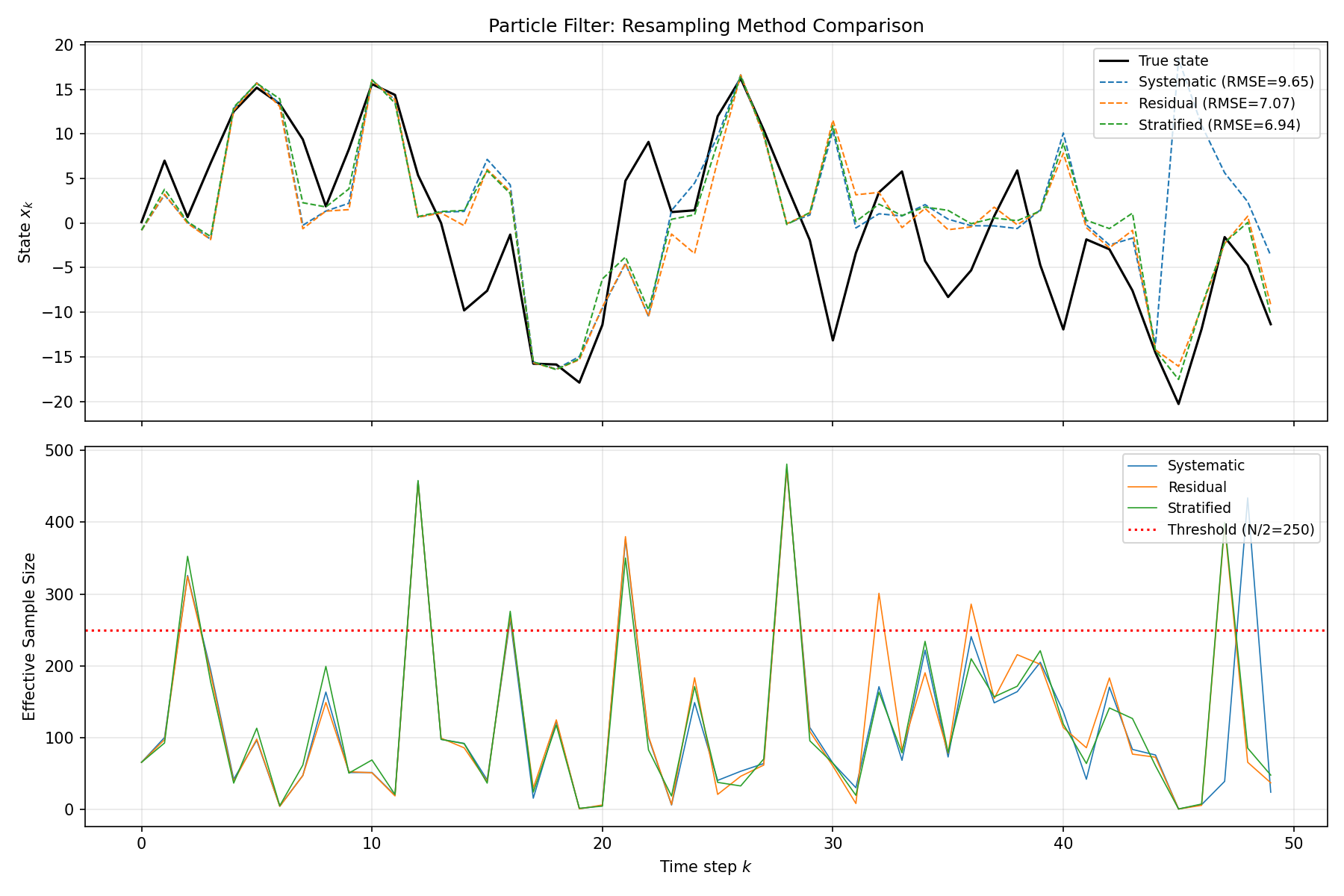
axes[0].set_title("Particle Filter: Resampling Method Comparison")
axes[0].legend(loc="upper right", fontsize=9)
axes[0].grid(True, alpha=0.3)
# ESS over time
for name, res in results.items():
axes[1].plot(res["ess"], color=colors[name], linewidth=0.8, label=name)
axes[1].axhline(y=N_particles / 2, color="red", linestyle=":",
label=f"Threshold (N/2={N_particles // 2})")
axes[1].set_xlabel("Time step $k$")
axes[1].set_ylabel("Effective Sample Size")
axes[1].legend(loc="upper right", fontsize=9)
axes[1].grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig("particle_filter_comparison.png", dpi=150)
plt.show()
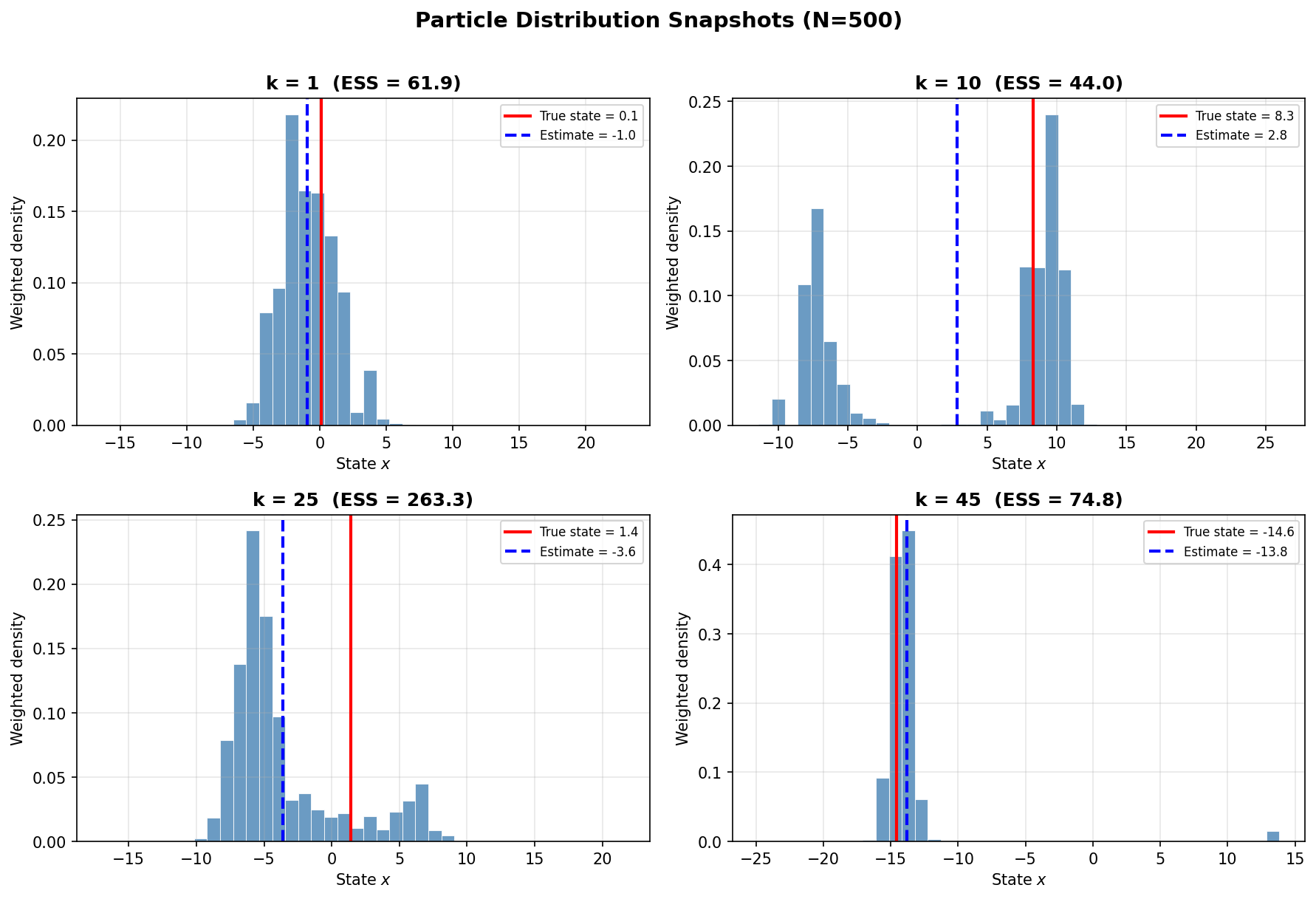
Particle Distribution Snapshots
The figure below shows the weighted particle distribution (histogram) at selected time steps. The red line indicates the true state, and the blue line shows the estimate.
Comparison of Filtering Methods
The following table compares the particle filter with Gaussian-approximation filters.
| Property | Particle Filter | UKF | CKF |
|---|---|---|---|
| Distribution assumption | None (arbitrary) | Gaussian | Gaussian |
| Computational cost | \(O(N)\) (depends on particle count) | \(O(n^3)\) | \(O(n^3)\) |
| Parameters | Particle count \(N\), ESS threshold | \(\alpha, \beta, \kappa\) | None |
| Multimodality | Supported | Not supported | Not supported |
| High dimensions | Difficult (curse of dimensionality) | Good | Good |
| Implementation complexity | Moderate | Moderate | Low |
The particle filter’s key advantage is its freedom from distributional assumptions, but it faces the “curse of dimensionality” in high-dimensional problems where the required number of particles grows exponentially. For low-dimensional problems with strong nonlinearity or non-Gaussianity, the particle filter is the method of choice. For high-dimensional problems with moderate nonlinearity, CKF or UKF are more practical.
Related Articles
- Kalman Filter: Theory and Python Implementation - The optimal filter for linear Gaussian systems.
- Extended Kalman Filter (EKF): Theory and Python Implementation - Linearization-based approach for nonlinear systems.
- Unscented Kalman Filter (UKF): Nonlinear State Estimation Theory and Python Implementation - Sigma-point based nonlinear filter.
- Kalman Smoother (RTS Smoother): Theory and Python Implementation - Smoothing using observations from all time steps.
- Markov Chain Monte Carlo (MCMC): Metropolis-Hastings and Gibbs Sampling - A Monte Carlo sampling method closely related to particle filters.
References
- Arulampalam, M. S., Maskell, S., Gordon, N., & Clapp, T. (2002). “A Tutorial on Particle Filters for Online Nonlinear/Non-Gaussian Bayesian Tracking.” IEEE Transactions on Signal Processing, 50(2), 174-188.
- Doucet, A., & Johansen, A. M. (2009). “A Tutorial on Particle Filtering and Smoothing: Fifteen Years Later.” Handbook of Nonlinear Filtering, 12(656-704), 3.
- Douc, R., & Cappe, O. (2005). “Comparison of Resampling Schemes for Particle Filtering.” Proceedings of the 4th International Symposium on Image and Signal Processing and Analysis, 64-69.